Blueprint-Free Life: Scientists Discover DNA Built Without Templates
A groundbreaking study published in Science (April 2026) has revealed a phenomenon that challenges a central dogma of biology. For decades, we believed that building DNA always required a “blueprint” (a template strand) for enzymes to copy. However, researchers at Stanford University have discovered a protein that can build specific DNA sequences entirely from scratch, using its own physical shape as the mold.
1. The Enzyme That “Self-Blueprints”
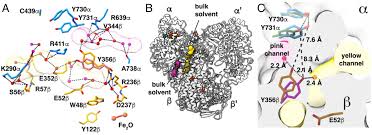
The discovery centers on a bacterial defense system called DRT3 (Defense-associated Reverse Transcriptase), specifically an enzyme known as Drt3b.
-
The Old Way: Typically, an enzyme called polymerase “reads” an existing DNA or RNA strand to create a matching new one.
-
The New Way: Drt3b doesn’t need to read anything. Its internal structure acts as a physical stencil. As it builds a DNA strand, the shape of the enzyme itself dictates which genetic “letters” (bases) are added, and in what order.
-
The Result: It produces a consistent, sequence-specific DNA strand without any external reference material. This is the first time “protein-templated” DNA synthesis has ever been observed in nature.
2. Why Bacteria Do This: The Viral Shield
This strange mechanism isn’t just a biological fluke; it’s a weapon.
-
Anti-Phage Defense: Bacteria use the DRT3 system to fight off phages (viruses that infect bacteria).
-
The Trap: By producing these specific, template-free DNA strands, the bacteria can interfere with the virus’s ability to replicate its own genetic code, essentially “jamming” the viral machinery before it can take over the cell.
3. Expanding the Genetic Alphabet
While one team discovered how DNA is built, another (reported in Science Advances, Jan 2026) has discovered a way to change what it is built from.
-
Six-Letter DNA: Scientists have successfully integrated “unnatural base pairs” (UBPs) into DNA nanostructures.
-
Fat & Skinny DNA: By adding a 5th and 6th letter (beyond the standard A, T, C, and G), researchers have created XenoAptamers. These can form “fat” or “skinny” DNA structures that are far more stable and chemically diverse than natural DNA.
4. Summary of the 2026 Genetic Revolution
| Feature | Traditional Understanding | 2026 Discovery |
| Synthesis | Requires a DNA/RNA Template | Protein-Templated (Template-Free) |
| Information Flow | DNA $\rightarrow$ RNA $\rightarrow$ Protein | Protein $\rightarrow$ DNA (Direct Sequence Control) |
| Complexity | 4-Letter Genetic Code | 6-Letter “Xeno” Code |
| Application | Basic Replication | Advanced Gene-Editing & Data Storage |